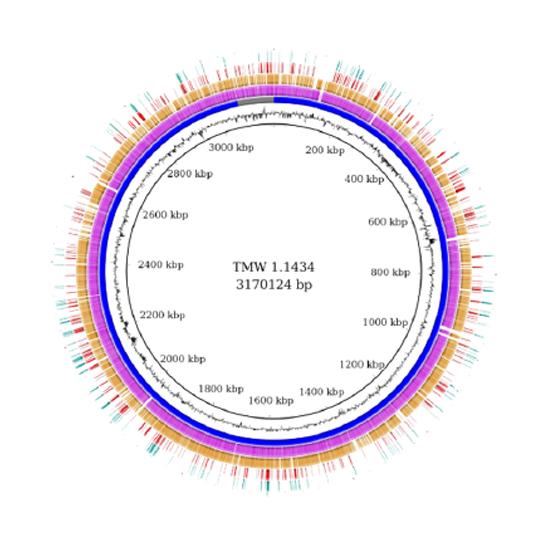
Quantitative Proteomics for the Comprehensive Analysis of Stress Responses of Lactobacillus paracasei subsp. paracasei F19
01-Sep-2017
J. Proteome Res., 2017, 16 (10), pp 3816–3829, DOI: 10.1021/acs.jproteome.7b00474
J. Proteome Res., online article
Lactic acid bacteria are broadly employed as starter cultures in the manufacture of foods. Upon technological preparation, they are confronted with drying stress that amalgamates numerous stress conditions resulting in losses of fitness and survival. To better understand and differentiate physiological stress responses, discover general and specific markers for the investigated stress conditions, and predict optimal preconditioning for starter cultures, we performed a comprehensive genomic and quantitative proteomic analysis of a commonly used model system, Lactobacillus paracasei subsp. paracasei TMW 1.1434 (isogenic with F19) under 11 typical stress conditions, including among others oxidative, osmotic, pH, and pressure stress. We identified and quantified >1900 proteins in triplicate analyses, representing 65% of all genes encoded in the genome. The identified genes were thoroughly annotated in terms of subcellular localization prediction and biological functions, suggesting unbiased and comprehensive proteome coverage. In total, 427 proteins were significantly differentially expressed in at least one condition. Most notably, our analysis suggests that optimal preconditioning toward drying was predicted to be alkaline and high-pressure stress preconditioning. Taken together, we believe the presented strategy may serve as a prototypic example for the analysis and utility of employing quantitative-mass-spectrometry-based proteomics to study bacterial physiology.